Quality on the Line: Spoilage Microbes in Milk?
Milk is pasteurized primarily as a food safety measure. A secondary benefit is that almost all bacteria are killed-including those that cause milk to spoil. However, some spoilage bacteria may survive pasteurization.

 Milk is pasteurized primarily as a food safety measure. A secondary benefit is that almost all bacteria are killed-including those that cause milk to spoil. However, some spoilage bacteria may survive pasteurization.
Milk is pasteurized primarily as a food safety measure. A secondary benefit is that almost all bacteria are killed-including those that cause milk to spoil. However, some spoilage bacteria may survive pasteurization.
Microbes are classified according to distinguishing characteristics, such as optimum temperature and oxygen requirements for growth. Another practical classification is based upon cell morphology (cocci, rods, etc.) and staining reaction (Gram-positive [G+] or Gram-negative [G-]). The latter is based upon the preferential retention of selected stains by the bacterial cells. Those that retain crystal violet (bluish-purple) are classified as G+ while those that are stained with safranin (pinkish-red) are classified as G-.
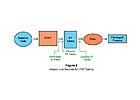
Many of the G+ bacteria are thermoduric (“heat enduring”) and capable of surviving, while most of the G- bacteria are psychrotrophic (capable of growth at refrigerated temperatures) and destroyed by pasteurization. Both types are present in raw milk. In addition, both G+ and the G- bacteria can produce heat-stable proteolytic and/or lipolytic enzymes that easily survive pasteurization. Raw milk containing high numbers of viable bacteria will contain high levels of liberated heat stable microbial enzymes. These enzymes may cause unclean, putrid, and bitter flavors in pasteurized milk that may have low viable bacteria counts. See Figure 1.
Psychrotrophic bacteria are the major cause of spoilage of HTST processed milk. The most common G- psychrotroph (capable of growth at refrigeration temperatures) in pasteurized milk is a member of the genus Pseudomonas. Psychrotrophic bacteria in pasteurized milk are the result of post-pasteurization contamination, due to unsanitary equipment and environmental conditions. The most common G+ rod-shaped bacteria in raw milk are members of the genera Bacillus and Clostridium. Some of these bacteria are both thermoduric (“heat enduring”) and psychrotrophic. They grow at a slower rate than the G-’s and their impact on shelf-life may not be apparent if the product is kept below 40°F or consumed in less than 12 days. These slower growing bacteria have become a significant cause of spoilage in fluid products as code dates have been extended. Due to their slower growth and/or extended lag time (8–10 days), they may go undetected by the seven-day Moseley Test.
 Trouble Shooting a Microbe Problem: Systematic testing of aseptic line samples from raw receiving to packaging is required for pinpointing possible sources of contamination. Identification of the type of bacteria involved is helpful in determining if they survived HTST processing or if post-pasteurization contamination has occurred.
Trouble Shooting a Microbe Problem: Systematic testing of aseptic line samples from raw receiving to packaging is required for pinpointing possible sources of contamination. Identification of the type of bacteria involved is helpful in determining if they survived HTST processing or if post-pasteurization contamination has occurred.
Bottom Line: Spoilage bacteria can survive conventional HTST pasteurization. Spore-forming G+ rods are the predominant survivors. Some are capable of growth at refrigerated temperatures and contributing to spoilage when code dates are extended. n
Join Randolph Associates in Murfreesboro, Tenn. for an in-depth HTST Workshop May 2007. For registration information, contact Kristy Clark at 205/595-6455 or kristy.clark@raiconsult.com.


Figure 1. Microbes and microbial enzymes in raw and pasturized milk.
Microbes are classified according to distinguishing characteristics, such as optimum temperature and oxygen requirements for growth. Another practical classification is based upon cell morphology (cocci, rods, etc.) and staining reaction (Gram-positive [G+] or Gram-negative [G-]). The latter is based upon the preferential retention of selected stains by the bacterial cells. Those that retain crystal violet (bluish-purple) are classified as G+ while those that are stained with safranin (pinkish-red) are classified as G-.
Many of the G+ bacteria are thermoduric (“heat enduring”) and capable of surviving, while most of the G- bacteria are psychrotrophic (capable of growth at refrigerated temperatures) and destroyed by pasteurization. Both types are present in raw milk. In addition, both G+ and the G- bacteria can produce heat-stable proteolytic and/or lipolytic enzymes that easily survive pasteurization. Raw milk containing high numbers of viable bacteria will contain high levels of liberated heat stable microbial enzymes. These enzymes may cause unclean, putrid, and bitter flavors in pasteurized milk that may have low viable bacteria counts. See Figure 1.
Psychrotrophic bacteria are the major cause of spoilage of HTST processed milk. The most common G- psychrotroph (capable of growth at refrigeration temperatures) in pasteurized milk is a member of the genus Pseudomonas. Psychrotrophic bacteria in pasteurized milk are the result of post-pasteurization contamination, due to unsanitary equipment and environmental conditions. The most common G+ rod-shaped bacteria in raw milk are members of the genera Bacillus and Clostridium. Some of these bacteria are both thermoduric (“heat enduring”) and psychrotrophic. They grow at a slower rate than the G-’s and their impact on shelf-life may not be apparent if the product is kept below 40°F or consumed in less than 12 days. These slower growing bacteria have become a significant cause of spoilage in fluid products as code dates have been extended. Due to their slower growth and/or extended lag time (8–10 days), they may go undetected by the seven-day Moseley Test.

Figure 2. Aseptic Line Samples for HR3 Testing
- Priority: Eliminate in-plant post-pasteurization contamination. This can be confirmed by STRESS (70°F for 18 – 20 hrs) coliform (Dairy Foods: November 2005) and Moseley (seven days at 45°F) or by HR1 and HR2 testing (Dairy Foods: March 2006). Once the presence of G- bacteria has been eliminated and shelf-life problems are still present-testing for G+ thermodurics should be initiated. Note: Testing for G+ bacteria when G- bacteria are present may yield conflicting and confusing results.
- Testing for G+’s: An excellent method to check for these bacteria is the HR-3 screening test (Dairy Foods: March 2006). It is designed for screening raw and pasteurized milks for the presence of G+, spore-forming rods. The protocol uses a vivid color indication for positive samples. It is simple to perform and yields timely results (usually within 14 hours). The presence of large G+ rods is confirmed by microscopic examination. See Figure 2 for suggested aseptic line sampling for HR3 testing.
- Reacting to HR3 Results: If G+ bacteria are present in large numbers at the HTST balance tank, aseptic samples from raw silos should be tested. If they are confirmed in raw silos, each incoming load of raw milk should be checked to determine loads/producers contributing to high Gram + numbers. Effective sanitation practices from the farm to the package are required to correct the problem.
Bottom Line: Spoilage bacteria can survive conventional HTST pasteurization. Spore-forming G+ rods are the predominant survivors. Some are capable of growth at refrigerated temperatures and contributing to spoilage when code dates are extended. n
Join Randolph Associates in Murfreesboro, Tenn. for an in-depth HTST Workshop May 2007. For registration information, contact Kristy Clark at 205/595-6455 or kristy.clark@raiconsult.com.
Looking for a reprint of this article?
From high-res PDFs to custom plaques, order your copy today!




